Clustered regularly interspaced short palindromic repeats (CRISPR) loci occur in the prokaryotic genome. These loci consist of DNA sequences acquired from foreign DNA and serve as a template for CRISPR guide RNA. CRISPR loci also encode proteins called CRISPR associated proteins (Cas). Some of these Cas proteins possess nuclease activity and play a vital part in the bacterial and archaeal defence system. Cas proteins are guided by guide RNA (gRNA) to target and degrade invading viral genomes and plasmids.
In 2005, Alexander Bolotin discovered the first Cas protein with nuclease activity while studying the genome of Streptococcus thermophilus. The protein he discovered is now known as Cas9. In 2017, Feng Zhang and his team demonstrated successful genome editing via the CRISPR-Cas9 system in eukaryotic cells. They engineered Cas9 orthologs isolated from Streptococcus pyogenes and Streptococcus thermophilus. At present, there are 287 reviewed and more than 250,000 putative Cas proteins. So far, only Cas9 and Cas12a nucleases show genome editing ability.
Applications of CRISPR/Cas
Applications of CRISPR/Cas based genome editing are:
- Identification and screening of functional genes and drug targets
- Generating engineered animal models of human diseases
- Correcting defective genotypes in the cells
- Developing sequence-specific anti-microbials and anti-virals
- Engineering crops to improve yield along with disease and drought tolerance
- Improving traits in livestock
- Engineering microorganisms for the production of bioplastics and biofuels
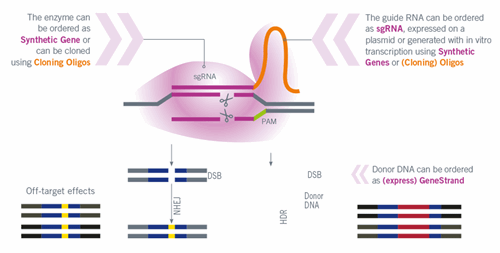
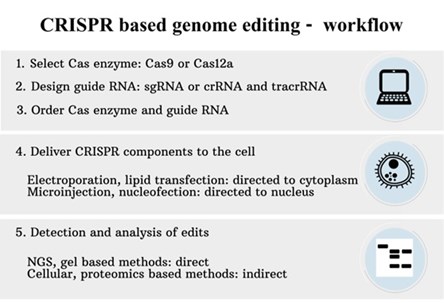
Workflow – CRISPR/Cas based genome editing
Experiment design
Choose an appropriate Cas enzyme (Cas9/Cas12a). The choice of Cas enzyme depends on the type of Protospacer Adjacent Motif (PAM) sequence. Cas enzyme requires PAM sequence to cleave the DNA.
After the selection of Cas enzyme, gRNA is designed. gRNA guides the Cas enzymes towards its site of action. In the case of Cas9, either single guide RNA (sgRNA) or 2 part guide RNA is used. The 2 part guide RNA comprises of a duplex of CRISPR RNA (crRNA) and trans-activating CRISPR RNA (tracrRNA). tracrRNA consists of constant sequence to which Cas9 binds. crRNA consists of a spacer region that directs Cas9 to its target. In contrast, Cas12a requires a single crRNA.
Delivery of CRISPR/Cas to the target cells
CRISPR/Cas components can be introduced to the cells in the form of DNA, RNA or as ribonucleoprotein (RNP) complex. Transcription or/and translation are required in case of DNA and RNA. Furthermore, after the synthesis of Cas in the cytoplasm, it has to cross the nuclear envelope to reach the target site. Therefore, a nuclear localisation signal (NLS) needs to be embedded in the coding sequence of Cas. For RNP complexes, there is no need for translation or transcription and they can be transfected directly to the nucleus.
For the transport of CRISPR/Cas components directly to the nucleus, microinjection or nucleofection methods are preferred, while for the delivery of components to the cytoplasm electroporation and lipofection are utilised.
Detection of edits and analysis
To confirm that target cells are edited correctly, various methods can be used. Commonly used methods are sequencing and gel-based visualisation. Sometimes, indirect methods of detection (cellular and proteomic methods) are also used. The selection of the method depends on experimental needs, time and budget. Next generation sequencing is the gold standard for the detection and analysis of edits, and provides high accuracy and sensitivity.

Did you know that Eurofins Genomics offers INVIEW CRISPR Check, a service to analyse your CRISPR-mediated gene editting, as well as a broad range of gene Expression Services such as transcriptome analysis?