Overview – Introduction to targeted sequencing
Our targeted resequencing services use high-throughput next-generation sequencing (NGS) for analysis of genomic regions of interest across large numbers of samples with short turnaround times. It is therefore ideal for population studies, discovery and confirmation of SNPs and InDels and for the detection of rare variants. Eurofins Genomics provides complete project management with standard sequencing or customised solutions and complementary data analysis.
Applications – Advantages of targeted sequencing
- Targeted sequencing is ideal for:
- Focusing on specific regions of interest
- Targeted analysis of microbial genes
- Detecting rare variants, sequence-focused content from multiple samples in parallel using multiplexing
- Achieving comprehensive coverage, yet generating a smaller, more reasonable amount of data
- Identifying causative variants like SNPs and InDels across multiple genomic regions
- Fast turnaround time for more samples
Workflow – Targeted sequencing methods & technologies
Targeted resequencing is a high-throughput NGS method used with increasing frequency for identifying causative mutations within target content in populations by isolating and sequencing target regions of the genome from a sample library.
Target enrichment
Sequencing enrichment targets genomic regions of interest with high specificity and sensitivity, enabling the detection of both rare and common variants.
Exome sequencing is likely the most commonly used targeted resequencing method. Exome sequencing enriches only the protein-coding or exon regions of the human genome, where most phenotype-causing variants are presumed to be. The utilised Agilent SureSelect Human All Exon V8 panel shines with improved enrichment and uniform coverage across target regions.
With exome sequencing, comprehensive coverage of coding regions can be achieved.
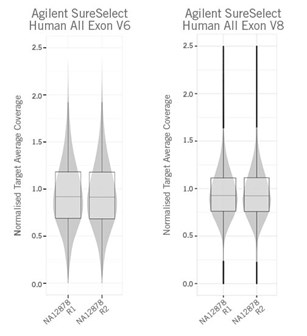
Comparison of normalised coverage distribution of Agilent SureSelect Human All Exon V6 and V8 panels. V8 panel with increased uniformity.
Gene-panel sequencing
Panel sequencing is a next-generation sequencing method for multiple genomic areas of interest. Panels can be pre-made or custom-designed to target cancer genes or DNA regions related to inherited diseases or specific phenotypes. By sequencing only genes of interest, gene panels greatly reduce sequencing costs, time and analytical effort.
Amplicon sequencing
Smaller target DNA regions can be amplified by PCR. Ultra-deep sequencing of the resulting PCR amplicons is a targeted approach usually used for applications like variant analysis and screening of somatic mutations from tumour samples.
Multiplexing options enable parallel high-throughput analysis of hundreds of genes from multiple samples at roughly the same time and cost as a comparable single-gene assay. This considerably facilitates variant discovery in populations and allows analysis of whole genes with their flanking regions and co-variations.
Eurofins Genomics offers a complete solution covering the entire process from experimental design, to high-fidelity PCR, NGS of amplicons and bioinformatics analysis for sequenced amplicons.
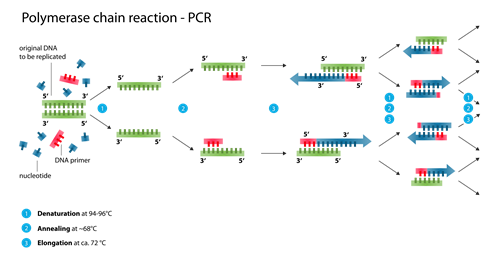
PCR amplification for Sanger sequencing
Target enrichment and capture using next-generation sequencing technologies with high-sequencing coverage is a popular method of targeted sequencing. Alternatively, for sequencing a small number of targeted regions, PCR amplification can be used in combination with Sanger sequencing. This method identifies base positions in dissimilar pairs of genes, SNPs or small insertions and deletions in genomic DNA sequences. This is commonly used to locate heterozygotes, detect genetic rearrangements, and uncover rare variants.
16S rRNA sequencing and ITS sequencing
Sequencing the hypervariable regions of the 16S ribosomal RNA gene or ITS region is a related amplicon sequencing method used to identify and compare bacteria or fungi present within a given sample. Studying complex microbiomes or environmental samples is simplified by options for pooling PCR amplicons together. Highly sensitive, full-length 16S rRNA gene amplicon sequencing permits accurate phylogenetic microbial characterisation and quantification down to genus or even species level.
